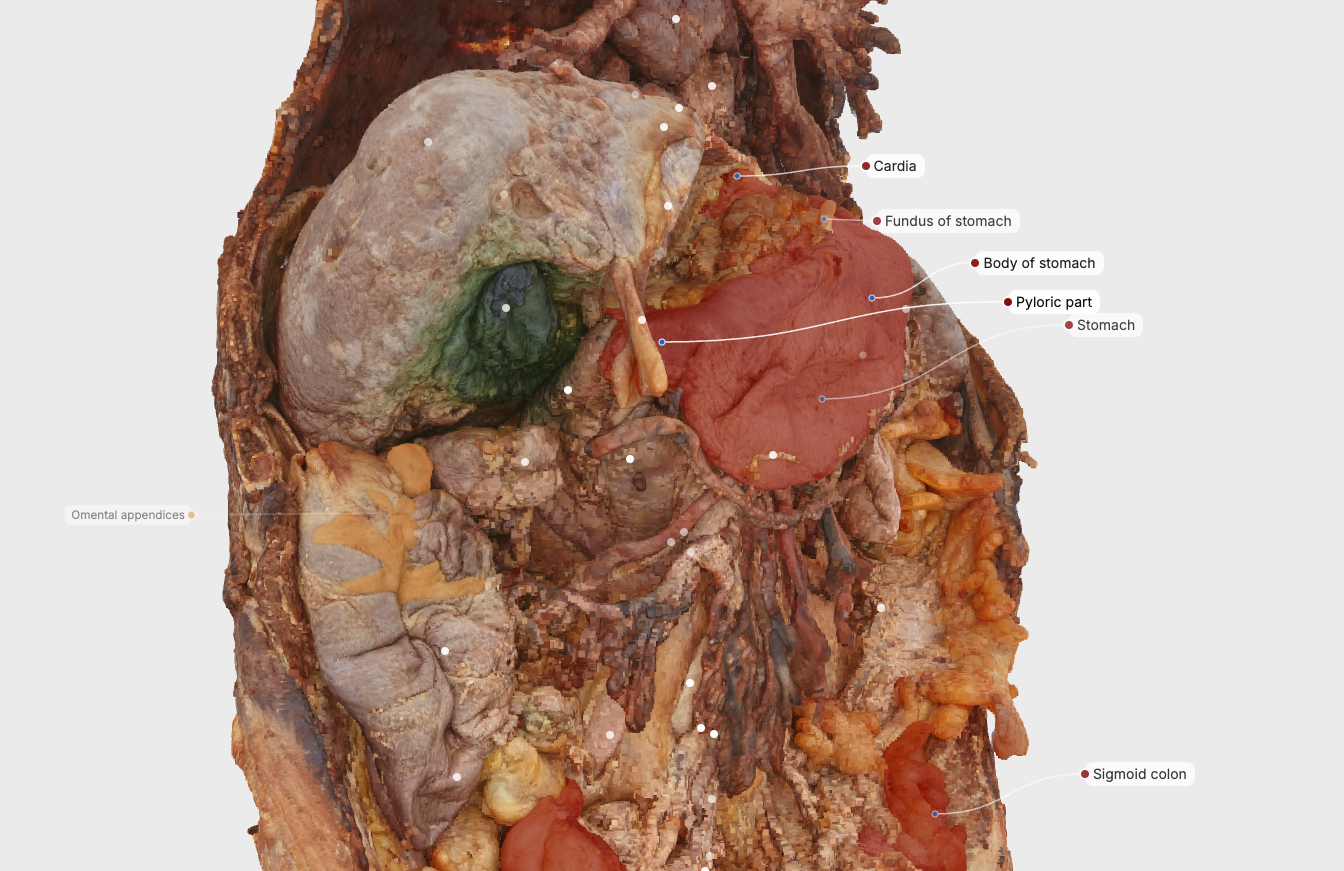
Introduction:
Graphics, spatial data and creating smart ways of combining data with interaction has always been a big interest and passion for me. As with an app like Enatom where these topics are prominent and ever evolving, it made sense to explore ways of visualizing and interacting with our data based on methods we have not already explored before. In this blog post I want to briefly talk about my experimentation and findings about signed distance fields and how these bring potential to visualizing anatomical data.
Pointclouds
Enatom renders anatomical content using point clouds rather than traditional triangle meshes.
Point-based representations allow us to work directly with high-resolution photogrammetry data from real donated bodies, preserving anatomical variation without aggressive surface smoothing. This makes them well suited for anatomical realism.
At the same time, point clouds introduce specific spatial challenges:
- Spatial continuity is implicit rather than explicitly defined
- Surface orientation must be inferred locally
- Dense and sparse regions behave differently under lighting and clipping
- Reasoning about inside versus outside of structures is non-trivial
These are not flaws, but trade-offs. Still, they raise questions about how to reason more continuously about anatomical space.
Signed distance fields as a spatial representation
A signed distance field is a scalar field defined over space. At every point, it stores the distance to the nearest surface:
- Negative values indicate points inside a structure
- Positive values indicate points outside
- Zero represents the surface
Instead of describing anatomy as discrete samples, a signed distance field describes it as a continuous spatial function. This makes it useful for spatial queries, proximity reasoning, and smooth transitions — capabilities that are difficult to derive directly from point data alone.
Deriving spatial information from the field
Signed distance fields encode more than surface location.
By sampling the field, gradients can be computed at arbitrary positions. These gradients provide stable surface normals, even in regions where point density varies. This can support more consistent lighting, reduced visual noise, and a stable appearance across zoom levels.
Distance values themselves can also be visualized, for example through depth-based gradients or proximity highlighting. Used carefully, these techniques can reinforce spatial relationships without overwhelming the scene.
Navigating anatomy using continuous space
Traditional clipping planes often produce abrupt cut-offs when navigating dense anatomical regions.
Distance fields make it possible to slice through anatomy more gradually, with smooth transitions that preserve surrounding context. Structures can fade in or out based on distance rather than binary visibility, reducing sudden loss of orientation during exploration.
Because the field is continuous, it also allows interpolation between states. Anatomical layers can transition smoothly while maintaining spatial relationships, rather than appearing or disappearing abruptly.
Depth cues and spatial orientation
Distance fields integrate naturally with depth-aware rendering techniques such as distance-based ambient occlusion or volumetric shading.
These effects are not about visual polish. Their primary value lies in reinforcing depth cues—helping users understand what lies anterior or posterior, superficial or deep—especially in complex regions where multiple structures overlap.
Geometry stabilization and spatial robustness
Because signed distance fields describe volume rather than discrete samples, they can help detect gaps or inconsistencies in point-based data.
They provide a more reliable basis for spatial queries even when point density varies, supporting smoother interaction and more predictable behavior across different anatomical regions.
Relation to volumetric imaging
There is a conceptual overlap between signed distance fields and volumetric datasets such as CT or MRI scans.
This raises research questions around converting segmented volumetric data into distance fields, and around combining surface anatomy with volumetric information. While exploratory, this direction could help bridge gross anatomy with clinical imaging contexts.
Interaction and annotation in dense anatomy
A field-based representation enables interactions that are spatial rather than purely geometric:
- Proximity-aware selection
- More reliable hit-testing
- Awareness of inside and outside for complex structures
One particularly promising application is annotation.
Signed distance fields allow the system to answer questions such as how far a label is from the nearest surface, whether it intersects anatomy, and in which direction it can be safely offset. This enables surface-aware label placement, collision-free annotations, and callouts that adapt smoothly to zoom and rotation.
The same spatial reasoning could support measurement tools, region-of-interest highlighting, and guided exploration paths.
Closing
Signed distance fields are fascinating to me. Their concept is simple and has been a known thing for ages and it excites me to figure out new ways of using this concept for visualizing and interacting with anatomical data within Enatom. I’m excited to see what’s in store for these concepts in the future.